By Thom Sanders:

Sailfin mollies, orange chromide, roundtail paradisefish—these are just a few of the colorful (and colorfully named) ornamental fish varieties potentially susceptible to megalocytiviruses (MCVs), a genus of viruses that can trigger die-offs in freshwater and marine fish populations.
These viruses are the focus of dissertation research conducted by Samantha Koda, a PhD student in ID&I. Samantha recently secured a 2018 Australia-Americas PhD Research Internship which will help support her megalocytivirus research at the University of Sydney. There, she will be working in the laboratory of Dr. Joy Becker to validate a quantitative PCR (qPCR) assay for MCVs.
Three strains of MCVs account for the majority of infections: Infectious spleen and kidney necrosis virus (ISKNV), red sea bream iridovirus (RSIV), and turbot reddish body iridovirus (TRBIV).
Despite what their names might suggest, these subgroups lack host specificity, meaning they can infect multiple species. Combined, the three varieties have been reported in upwards of 125 species. Samantha notes that “the ability of this virus to infect so many species around the world has largely been attributed to the global ornamental and food fish trade.”
The viruses are especially devastating in the aquaculture industry where infections can cause mortality rates approaching 100%.
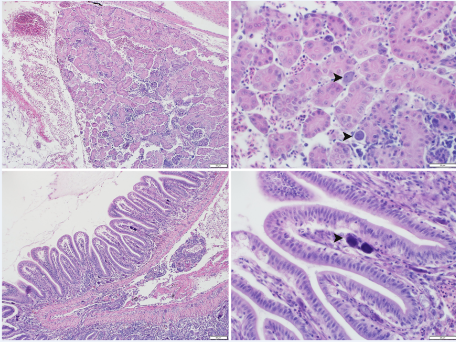
Unfortunately for researchers and fish health professionals, infected fish don’t exhibit telltale symptoms that might indicate an MCV infection. “When these fish are infected with MCV, behaviorally we see very typical signs of disease such as lethargy, abnormal position in the water column, splenomegaly, and increased gilling,” Samantha Koda says. “There are no gross clinical signs that are specific for megalocytivirus.”
Because of this, and because MCV infections can occur concurrently with bacterial infections, parasites, fungi, and other viruses, MCVs are tricky to diagnose. This makes it difficult for scientists and fish health professionals to determine if MCVs are responsible when symptoms appear in fish populations.
But Samantha’s research aims to change that.
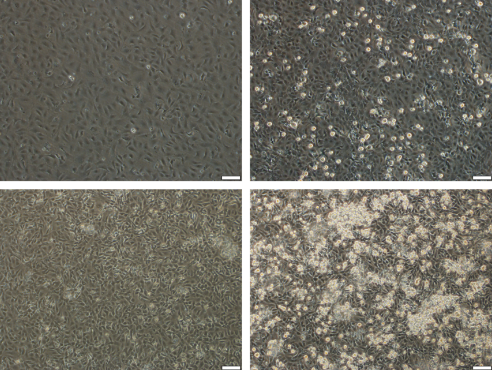

Her current goal is to validate a real-time qPCR assay that targets the three main strains of MCV. This tool would help researchers and fish health professionals quickly and confidently identify MCV infections in fish populations.
In addition to developing molecular diagnostics, Samantha’s research focuses on the genomic characterization of MCVs. She believes that characterizing the viruses’ genomes could help scientists better understand how this virus is able to infect such a large number of species. Using genomic characterization, Samantha says, “We can now look at certain genes that this virus carries or even acquired from their hosts and improve our knowledge on its possible virulence factors.”
Samantha hopes that MCV research will eventually lead to a vaccine for ornamental fish.
For more information about megalocytiviruses and their impact on ornamental fish aquaculture, check out the related IFAS Extension publication written in part by Dr. Thomas Waltzek, an Assistant Professor in ID&I.





